
A cluster of 35 FPGA works both as testbed for the neuroAIˣ framework and as fully functional neuroscience simulators.
Despite decades of research, the brain remains to a large extent a mystery. There has been tremendous progress in understanding the building blocks of the brain – neurons and synapses – and the delicate interplay between different regions of the brain to support cognitive functions. However, the basic question of how the brain processes information is still a puzzle. This is a fundamental question not only for neuroscience and medicine, but also for engineers and computer scientists, who are increasingly taking inspiration from the brain to improve the architecture and performance of computers.
To answer this question, neuroscientists look at groups of neurons – so-called microcircuits – and at the interplay of small brain areas. This level of analysis allows to study how individual neurons work together to form circuits and to perform complex tasks. It allows neuroscientists to develop models to explain how the brain processes information and, in the longer run, how behavior emerges from the activity of the neurons. This type of investigation relies heavily on computer simulations of biological neural network models.
“As a neuroscientist I am excited that Gemmeke and his team address the hard problem of how to cope with the immense number of contact points of each individual neuron straight on. Instead of using a toy model for demonstration, they employ a neuronal network model capturing all the neurons and all the synapses within a cubic millimeter of brain tissue. The progress reported here is breathtaking. Simulations much faster than realtime are essential for investigations of plasticity and learning unfolding over hours and days. But more importantly, the approach enables the team to systematically determine the bottlenecks on the way to a generic neuromorphic computing system.” says Markus Diesmann, director and neuroscientist at the FZ Jülich.
Today, biological neural networks models are simulated using either specialized software on high-performance compute clusters, or customized hardware systems tailored for this type of applications. The simulations allow us to test and refine our understanding of how biological neural networks work. New insights, however, often result in new requirements on the assumptions of the model and on the specifications of the simulator. While software can flexibly accommodate these changes, customized hardware systems face a chicken-and-egg problem: the better the hardware is fitted to a certain model, the faster it will generate new insights that will in-turn triggers a revision of the hardware itself.
From the perspective of a system engineer, this chicken-and-egg problem is a nightmare. “When I started my PhD in the field of neuromorphic computing, I wanted to construct a digital machine that is good at simulating biological neural networks like those in the human neo-cortex,” explains Kevin Kauth, PhD student at RWTH Aachen University. “However, it became quickly apparent that there is a major lack of ‘specifications’ of biological operating principles and of universally accepted benchmarks for validation and test. The focus of our team then shifted from targeting an architecture optimized for a specific application, to developing a framework suited to narrow down a fuzzy and moving target.”
The framework developed by Kauth and colleagues is composed by three pillars. The first pillar addresses the problem of simulating with digital hardware the massive interconnect network of the brain – the connectome. This is a far from trivial problem: a neuron collects inputs from and sends output to many other neurons — on the order of 10,000 on average for both input and output for a mammalian neuron. By contrast, a transistor has only three nodes for input and output all together. To build a simulator it is necessary to make drastic choices on the type of hardware employed and on the connectivity between the different computing nodes, and these choices have a major impact on the efficiency of the simulator itself. To trim out unsuitable directions, the group has developed a fast simulation tool that allows to explore within seconds the capabilities of networks of thousands to millions of compute nodes. This software tool, dubbed the static simulator, is based on much simplified assumptions of the network and does not account for bottlenecks such as memory and compute latency.
These bottlenecks are fully taken into account in the second pillar of the neuroAIˣ framework: the dynamic simulator. This is a virtual prototyping tool that allows to model the behavior of the customized hardware systems, incorporating dynamic effects such as congestion. Architectural components like memories, routers and schedulers are described in C++ in a modular and hierarchical way, which allows to flexibly explore various scenarios in terms of hardware choices and connectivity. As much as it provides precision, it stays behind in performance.
The third pillar of the neuroAIˣ framework is a cluster formed by 35 field-programmable gate array (FPGA) boards. The cluster, which has been realized following the insights given by the static and dynamic simulators, serves in first place as a hardware platform to calibrate the dynamic simulator itself (i.e., to extract empirical information on parameters such as the time needed to compute the dynamics of a single neuron or the memory bandwidth utilization) and to verify its accuracy. After calibration, the dynamic simulator is able to make precise quantitative prediction on the behavior of the system.

A cluster of 35 FPGA works both as testbed for the neuroAIˣ framework and as fully functional neuroscience simulators.
A field-programmable gate array (FPGA) is an integrated circuit designed to be configured by a customer or a designer after manufacturing. An FPGA contains an array of programmable logic blocks and memory elements, and a hierarchy of reconfigurable interconnects that allows the blocks to be wired together.
“Having a reliable digital twin is crucial for the development of next-generation neuroscience simulators,” explains Prof. Tobias Gemmeke, head of the Chair of Integrated Digital Systems and Circuit Design (IDS) at RWTH, and supervisor of the whole project. “It allows us to study in a very flexible way the effects of different network topologies or what happens if we scale up the system, without the burdens of developing new hardware. At the same time, the FPGA cluster is itself a very flexible hardware platform, suitable for further explorations. For instance, to implement a different network topology, we can change the cabling of the cards or just modify the settings. To study different neuron models, we can reprogram the FPGAs.”
The FPGA cluster is however more than just a training set for the neuroAIˣ framework: it is itself a fully functional neuroscience simulator, which surpass the best platforms available today both in terms of speed and energy efficiency. To test the capabilities of the FPGA cluster, Kauth and colleagues have chosen the so-called microcircuit model. With its 77.169 neurons and 300 million synapsis, the microcircuit model represents 1 mm2 of the sensory cortex of early mammalian and is one of the smallest models that exhibits a behavior that matches in-vivo observations.
On the neuroAIˣ FPGA cluster, simulations of the microcircuit model run 20 times faster than the biological real-time (BRT) and almost 10 times faster than on the fastest simulators available today. Such an acceleration is important for gaining insights on neurological processes that occur over long time-scales, such as learning.
“We were pleasantly surprised by the high acceleration achieved by our system,” says Kauth, “because the key focus of our work was on flexibility and reproducibility of the simulator system. We can say, we killed two birds with one stone!”
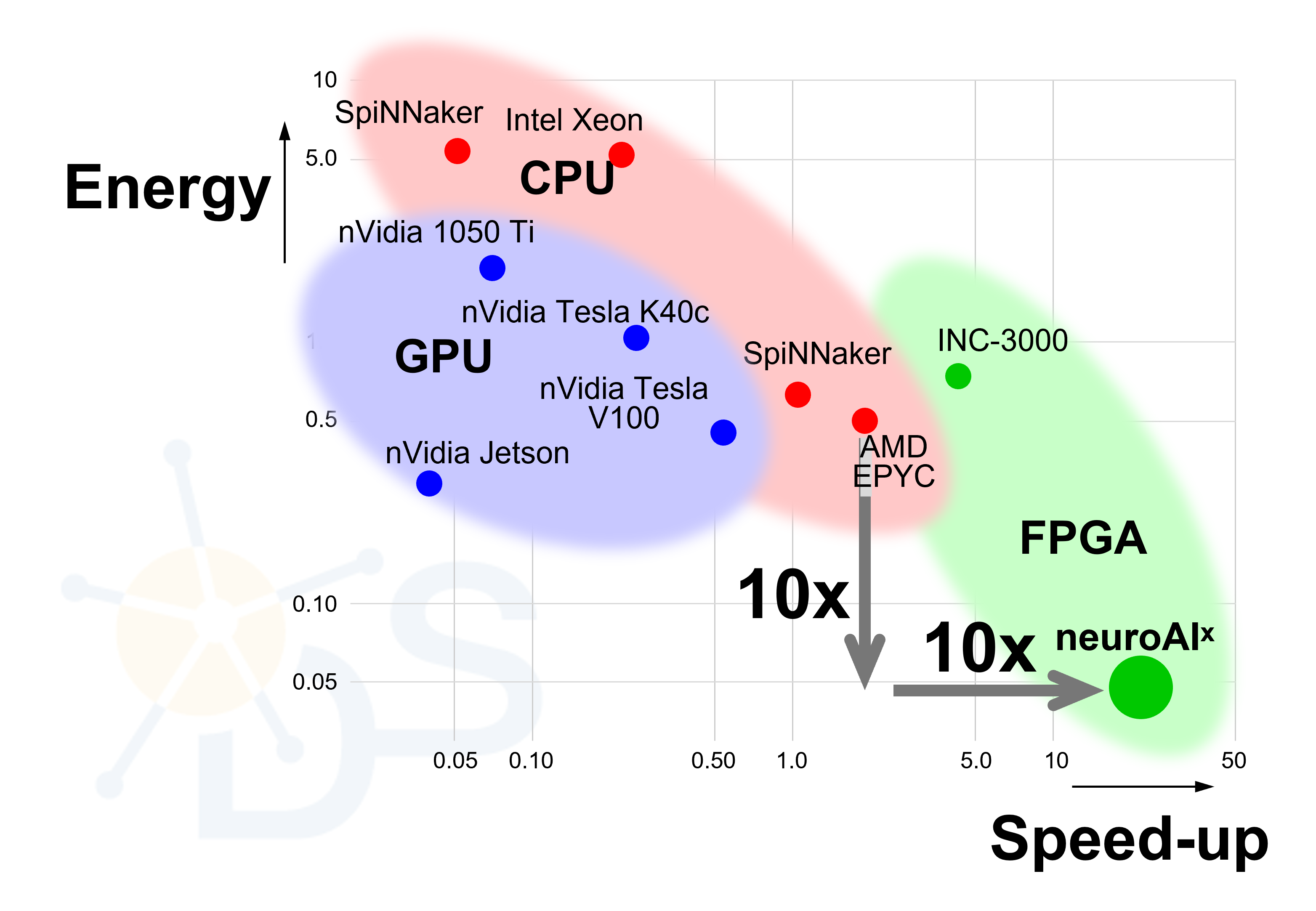
neuroAIˣ FPGA cluster is ten time faster and ten time more energy efficient than today’s best neuroscience simulators at running of biological neural networks.
The FPGA cluster is not only faster than other neuromorphic simulators, but it is also more energy efficient – again a factor 10 more efficient than today’s best platforms. Efficiency and speed go hand in hand, and both result from the fact that the FPGA cluster has been designed exploiting known neuromorphic principles.
“The capabilities of modern-day artificial intelligence (AI) are shooting through the roof – as is the energy consumed in computing hardware. Humanity badly needs to take inspiration from biology to conceive sustainable ways of realizing ‘smart’ computations,” says Gemmeke. Even if the microcircuit model used to benchmark the cluster is of little relevance for real-life applications, the high energy efficiency holds good promises that bio-inspired computing architecture can help mitigating the carbon footprint of future AI.
“What we achieved is just the first step towards more efficient hardware for data intensive applications,” adds Gemmeke. “To get anywhere close to the incredible efficiency of the brain at any cognitive task, we need a better understanding of how the brain transform information. Our FPGA cluster can play some role in this quest, as it is a platform where neuroscientists can run their simulations ten times faster than what is possible today.”
Furthermore, the FPGA cluster can be used to explore neuromorphic hardware architectures based on novel memristive devices, such as those researched in the projects Neurotec and NeuroSys – two major initiatives funded by the German Ministry of Education and Research (BMBF) for developing next-generation neuromorphic hardware. Memristors (the name is a contraction of “memory resistor”) are passive circuit elements whose resistance can be programmed by applying an external voltage. This feature makes them ideal candidates to develop the hardware-analog of synapses, and holds great promises for boosting the performance of neuromorphic hardware. The flexibility of the neuroAIˣ framework can facilitate the co-design of algorithms and hardware based on these novel devices by emulating the behavior of novel electronic devices on the FPGAs.
Gemmeke and colleagues would love to realize an upscaled version of the FPGA cluster and add a web interface to provide cloud-access to the cluster for neuroscientists and AI researchers around the world. This can be an attractive intermediate solution for neuroscience simulations until the next generation of truly neuromorphic platforms become available. “FPGAs are not the ultimate hardware solutions to implement biological neural networks, but their flexibility can accelerate the development of the whole field and support research on how biological neural networks can be used to solve real-world problems,” says Gemmeke.
The success of this endeavor – like many others – is pending on whether his team can activate sufficient human and financial resources for their research. The team is actively looking for donations to get it of the ground. Gemmeke explains: “While there is tremendous inertia in industry and investment by venture capital to apply AI, we see a huge interest from the general public, which is not only fascinated by the potential of AI, but also worried about its socio-economic consequences and its enormous carbon footprint. With a crowd-funding campaign everyone is invited to establish an alternative, i.e. sustainable, way of truly brain-inspired computing.” Updates will be made available here. A detailed description of the neuroAIx framework has been reported in the open-access journal Frontiers in Computational Neuroscience.
Free AI Website Creator